Mycetoma Genomics
Unlocking the Genetic Secrets of Madurella species.
Bridging the Gap Between Genomics, Identification, and Patient Care
Background
The Genomic Frontier: Understanding Fungal Pathogenesis
The transition from traditional microbiology to the genomic era has revolutionised our understanding of pathogenic fungi. For the genus Madurella, whole-genome sequencing (WGS) is not just a data exercise; it is a roadmap to survival.
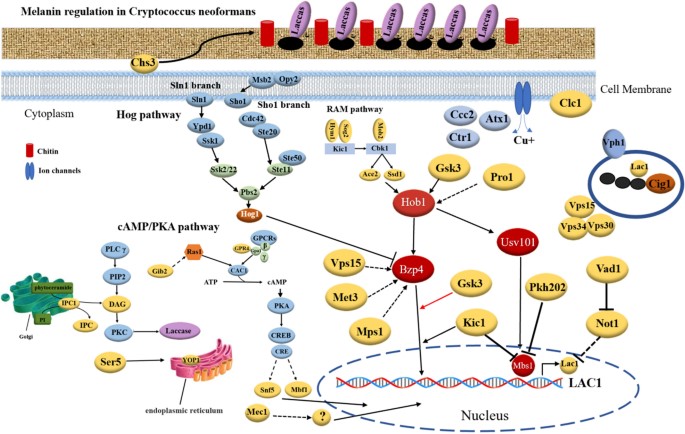
- Mapping the “Arsenal”: Genome data allows researchers to identify specific gene clusters responsible for melanin production, chitin synthesis, and grain formation. These are the primary mechanisms Madurella uses to shield itself from the human immune system and antifungal drugs.
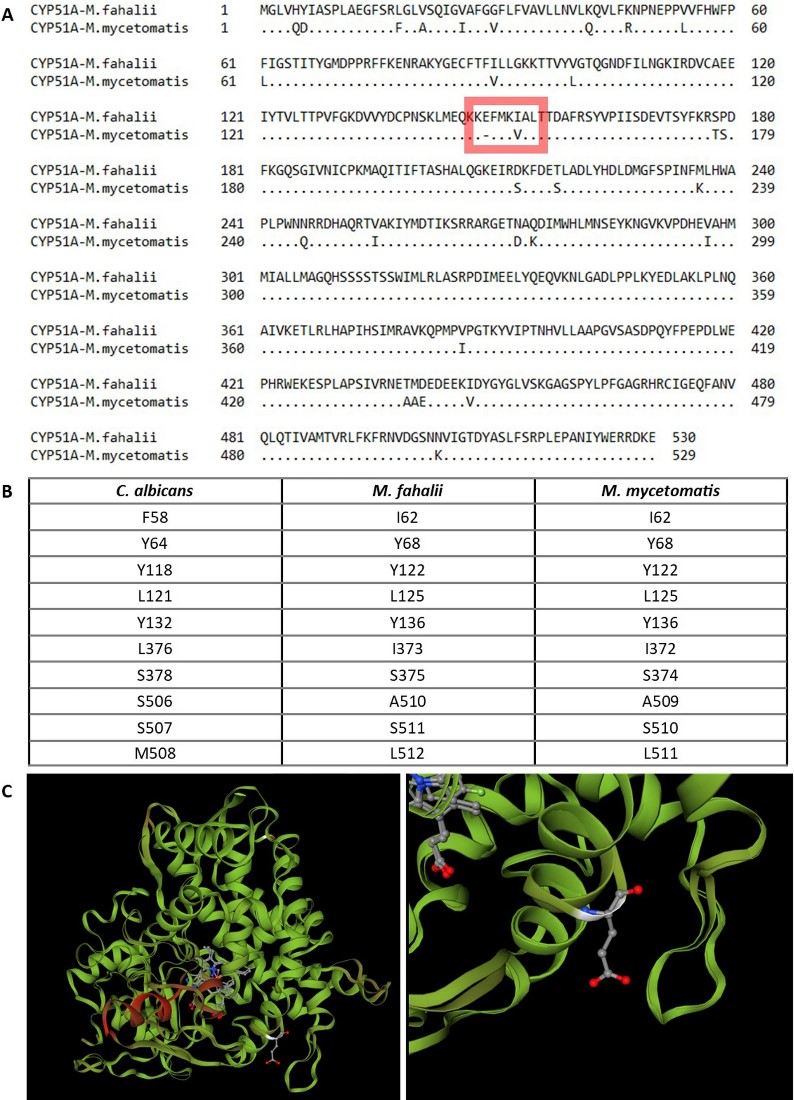
- Predicting Resistance: By analysing the Cytochrome P450 genes and other metabolic pathways, we can predict inherent resistance to common azoles, shifting treatment from “trial and error” to precision medicine.
- Host-Pathogen Interactions: Genomics helps us understand how these fungi adapt to the low-oxygen, high-stress environment of a human tissue, revealing new targets for therapeutic intervention.
The Precision of DNA Barcoding
Morphological identification of Madurella is notoriously difficult. Many species are “sterile” in culture (failing to produce spores) or belong to cryptic species complexes where different species look identical under a microscope but respond differently to treatment.
Why DNA Barcoding is the Gold Standard:
- Resolving Species Complexes: DNA barcoding, specifically targeting the ITS region, can distinguish between M. mycetomatis, M. fahalii, and M. tropicana, which are morphologically indistinguishable.
- Speed and Accuracy: While fungal cultures can take 3–4 weeks to grow, a DNA barcode can be sequenced and matched against a database in 48 hours.
- Universal Consistency: Unlike morphological traits that change based on growth media or temperature, the genetic “fingerprint” remains constant.

Representative Accession Numbers of Madurella species:
Click the Accession Numbers below to view the records directly in the NCBI database.
- Key Genome Assemblies (WGS)
| Species | Assembly Accession | Value to Research |
| M. mycetomatis | GCA_001275765.2 | The reference scaffold for grain-forming pathogens. |
| M. fahalii | GCA_003014105.1 | Essential for comparative studies in species diversity. |
| M. tropicana | GCA_003015055.1 | Critical for understanding regional variations in mycetoma. |
- Primary Diagnostic Barcodes (ITS)
| Species | Barcode Accession | Utility |
| M. mycetomatis | AY957291 | Primary reference for clinical PCR identification. |
| M. pseudomycetomatis | JF938052 | Discriminating from other black-grain causing agents. |
- Direct NCBI Genome Assembly Links
These represent the most complete genetic data available.
- GCA_001275765.2 (Madurella mycetomatis mm55) https://www.ncbi.nlm.nih.gov/datasets/genome/GCA_001275765.2/
- GCA_003014155.1 (Madurella mycetomatis v873) https://www.ncbi.nlm.nih.gov/datasets/genome/GCA_003014155.1/
- GCA_000507305.1 (Madurella mycetomatis CBS 109801) https://www.ncbi.nlm.nih.gov/datasets/genome/GCA_000507305.1/
- GCA_003014105.1 (Madurella fahalii CBS 129176) https://www.ncbi.nlm.nih.gov/datasets/genome/GCA_003014105.1/
- GCA_003015095.1 (Madurella pseudomycetomatis IFM 46460) https://www.ncbi.nlm.nih.gov/datasets/genome/GCA_003015095.1/
- GCA_003015055.1 (Madurella tropicana CBS 201.38) https://www.ncbi.nlm.nih.gov/datasets/genome/GCA_003015055.1/
Data Synergy: BioProjects
For large-scale analysis, these BioProjects aggregate raw sequencing data (SRA), samples, and assemblies into a single research node:
- PRJNA212351: The primary initiative for Madurella genomic architecture.
- PRJNA782605: A global effort to map the diversity of mycetoma-causing agents.
Specialised Fungal ITS & Barcoding Databases
While GenBank offers a vast volume of data, the following platforms provide curated “Gold Standard” sequences essential for accurate identification of species complexes like Madurella.
- UNITE: The Unified System for Fungal Identification
UNITE is the world’s leading database for the molecular identification of fungi. It uses a unique “Species Hypothesis” (SH) system to cluster sequences based on similarity thresholds.
- Focus: All fungi, with heavy emphasis on environmental and clinical isolates.
- Key Feature: Provides Digital Object Identifiers (DOIs) for species hypotheses, allowing researchers to cite specific genetic clusters even if the species is not yet formally named.
- Link: UNITE Database
- ISHAM-ITS: Medical Mycology Reference Database
Managed by the International Society for Human and Animal Mycology, this is the “clinical gold standard.” Every sequence in this database is quality-controlled and derived from strains identified by experts.
- Focus: Human and animal pathogenic fungi (including Madurella).
- Key Feature: Ideal for clinical labs requiring high-confidence identification for diagnostic purposes. It avoids the “taxonomic noise” often found in larger, uncurated databases.
- Link: ISHAM Barcoding Database
- NCBI RefSeq Fungal Targeted Loci
A curated subset of the main NCBI database. Sequences here are primarily derived from type specimens (the original specimen used to describe the species).
- Focus: Taxonomically verified reference sequences.
- Key Feature: Accessions start with NR_ (e.g., NR_111107). These are the most reliable sequences for verifying if a specimen belongs to a specific genus or species.
- Link: NCBI RefSeq Fungi
- BOLD Systems (Barcode of Life Data System)
BOLD is a global informatics workbench that supports the acquisition, storage, and analysis of DNA barcodes.
- Focus: Broad biodiversity across all kingdoms, but with a robust fungal section.
- Key Feature: Integrates molecular data with morphological images and geospatial GPS data, providing a holistic view of where and how a species was discovered.
- Link: BOLD Systems
- MycoBank
While primarily a nomenclatural database, MycoBank links fungal names to their original descriptions and, increasingly, to their DNA barcodes.
- Focus: Fungal nomenclature and taxonomy.
- Key Feature: Use this to check if a Madurella species name has been recently updated or moved to a different genus (synonymy).
- Link: MycoBank
Summary Table for Webpage Sidebar
| Database | Best For… | Type of Data |
| UNITE | Environmental & General Research | Species Hypotheses (SH) |
| ISHAM | Clinical & Medical Diagnosis | Expert-verified Pathogens |
| RefSeq | Formal Taxonomy | Type Specimen Sequences |
| BOLD | Global Biodiversity | Barcodes + Images + Maps |
Design Tips for the Webpage:
- Call to Action: Include a “Download List” button that links to a curated text file of these accessions.
- Visual Hierarchy: Use the tables provided above; they are easier for researchers to scan than paragraphs of text.
- Interactive Links: Ensure every accession number is a direct hyperlink to NCBI. Researchers appreciate not having to copy-paste.
